Chinese researchers have developed interfacially polymerized porous polymer particles for low- abundance glycopeptide separation. These polymer particles – with hydrophilic-hydrophobic heterostructured nanopores – can separate low-abundance glycopeptides from complex biological samples with high-abundance background molecules efficiently.
Results were published in the journal Advanced Materials in an article entitled "Interfacially Polymerized Particles with Heterostructured Nanopores for Glycopeptide Separation."
Qualitative/quantitative analyses of low-abundance biomolecules from complex biofluids are critical in clinical diagnosis and prognosis. For example, glycosylated Aβ peptide can help identify Alzheimer's disease and circulating tumor DNA can help identify cancer.
Porous polymer materials have been extensively used for separation. However, most existing porous polymer materials have homogeneous compositions or pores. As a result, efficiently and specifically separating subsets of low-abundance biomolecules from complex samples (such as serum and plasma) is a great challenge. Although recent molecule-level surface modification efforts for these homogeneous porous polymer materials have demonstrated enhanced separation specificity, unspecific adsorption of high-abundance background molecules still exists.
Endogenous glycopeptides are significant biomarkers for many diseases, such as Alzheimer's disease and cancer. However, their separation is always plagued by the low abundance of glycopeptides (generally 10 to 500 pg/mL in biofluids) and the high abundance of background molecules (such as proteins and non-glycopeptides) in complex biofluids.
Researchers from the Technical Institute of Physics and Chemistry (TIPC) of the Chinese Academy of Sciences recently developed an emulsion interfacial polymerization approach to synthesize polymer particles with hydrophilic-hydrophobic heterostructured surfaces and two-dimensional Janus film actuators.
On the basis of previous studies, the researchers recently synthesized a series of polymer particles with hydrophilic-hydrophobic heterostructured nanopores by emulsion interfacial polymerization, realizing efficient separation of low-abundance glycopeptides from high-abundance proteins and non-glycopeptides.
The efficient separation is achieved by allowing solvent-dependent local adsorption of biomolecules on the heterostructured surface inside the nanopores via switching the solvent polarity.

Fig. 1. Design of materials with hydrophilic-hydrophobic heterostructured nanopores for solvent-switched molecule adsorption. a) Conventional hydrophobic porous materials. b) Conventional hydrophilic porous materials. c) Porous materials with hydrophilic-hydrophobic heterostructured nanopores. (Image by Dr. SONG Yongyang)
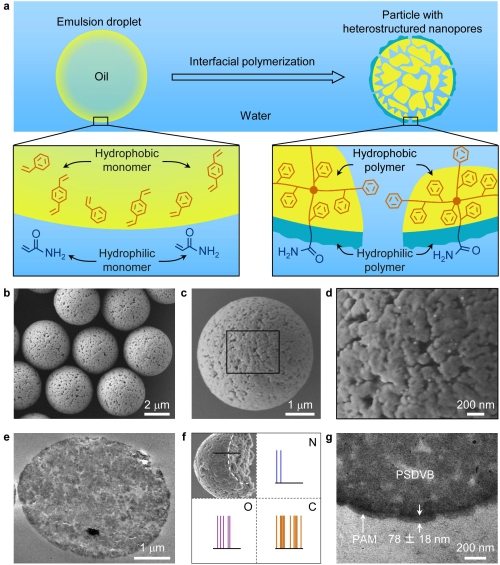
Fig. 2. Interfacially polymerized particles with hydrophilic-hydrophobic heterostructured nanopores. (Image by Dr. SONG Yongyang)
The researchers found that the particles with large nanopores (average pore size, 33 nm) can adsorb proteins and non-glycopeptides in high-polarity solvent via hydrophobic interactions, while the particles with small nanopores (average pore size, 3 nm) can adsorb glycopeptides in low-polarity solvent via hydrophilic interactions.
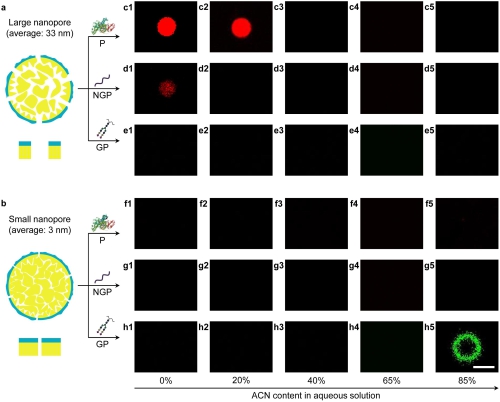
Fig. 3. Solvent-switched biomolecule adsorption by the particles with heterostructured nanopores. (Image by Dr. SONG Yongyang)
Thus, a two-step protocol was rationally designed to separate low-abundance glycopeptides from high-abundance proteins and non-glycopeptides. First, the hydrophobic region of the particles with large nanopores removed high-abundance hydrophobic proteins and non-glycopeptides in high-polarity solvent. Second, low-abundance hydrophilic glycopeptides were efficiently separated via the hydrophilic region of the particles with small nanopores in low-polarity solvent. The separation efficiency of the interfacially polymerized particles is superior to existing porous materials used for glycopeptide separation.
These particles provide a good candidate for application in the separation of low-abundance biomolecules from biological samples and downstream clinical diagnostics.
This work was financially supported by the National Science Fund for Distinguished Young Scholars and the Beijing Municipal Science and Technology Commission, among others.

